Projects
Vision Language Models for radiology and pathology
A visual-language model (VLM) pre-trained on natural images and text pairs poses a significant barrier when applied to medical contexts due to domain shift. Yet, adapting or fine-tuning these VLMs for medical use presents considerable hurdles, including domain misalignment, limited access to extensive datasets, and high-class imbalances. Hence, there is a pressing need for strategies to effectively adapt these VLMs to the medical domain, as such adaptations would prove immensely valuable in healthcare applications. We propose frameworks designed to adeptly tailor VLMs to the medical domain, employing knowledge grounding, selective sampling and hard-negative mining techniques for enhanced performance in retrieval tasks.

- Khan, Aisha Urooj, John Garrett, Tyler Bradshaw, Lonie Salkowski, Jiwoong Jason Jeong, Amara Tariq, and Imon Banerjee. “Knowledge-grounded Adaptation Strategy for Vision-language Models: Building Unique Case-set for Screening Mammograms for Residents Training.” MICCAI 2024 arXiv preprint arXiv:2405.19675 (2024).
- Chao, Chieh-Ju, Yunqi Richard Gu, Wasan Kumar, Tiange Xiang, Lalith Appari, Justin Wu, Juan M. Farina et al. “Foundation versus domain-specific models for left ventricular segmentation on cardiac ultrasound.” npj Digital Medicine 8, no. 1 (2025): 341.
- Ayoub, Chadi, Lalith Appari, Milagros Pereyra, Juan M. Farina, Chieh-Ju Chao, Isabel G. Scalia, Ahmed K. Mahmoud et al. “Multimodal fusion artificial intelligence model to predict risk for MACE and myocarditis in cancer patients receiving immune checkpoint inhibitor therapy.” JACC: Advances 4, no. 1 (2025): 101435.
- MICCAI2024, Deep-Breath workshop
- Ghosh, Shantanu, Vedant Parthesh Joshi, Rayan Syed, Aya Kassem, Abhishek Varshney, Payel Basak, Weicheng Dai et al. “Mammo-FM: Breast-specific foundational model for Integrated Mammographic Diagnosis, Prognosis, and Reporting.” arXiv preprint arXiv:2512.00198 (2025).
Large Language Models for clinical and biomedical domains
Domain adaptation – Generic large language models (LLMs), such as GPT-4, are trained on diverse datasets to develop a broad understanding of human language. While powerful across general domains, these models often struggle with the complexity and precision required in specialized fields like clinical medicine. Our project addresses this gap by developing domain-tailored LLMs specifically optimized for the clinical context of prostate and breast cancer.
Risk assessment and mitigation – Despite their transformative impact on natural language processing, public LLMs present notable vulnerabilities given the source of training data is often web-based or crowdsourced, and hence can be manipulated by perpetrators. We delve into the vulnerabilities of clinical LLMs, which is trained on publicly available biomedical literature and clinical notes, in the realm of data poisoning and model editing attacks. We are developing open-source toolkit for identification and recovery of the poisoned LLM.

- Tariq, Amara, Man Luo, Aisha Urooj, Avisha Das, Jiwoong Jeong, Shubham Trivedi, Haidar Abdul-Muhsin et al. “Development of LLM for Prostate Cancer—The Need for Domain-Tailored Training.” Cancer Detection and Diagnosis: A Handbook of Emerging Technologies (2025): 397.
- Das, Avisha, Ish A. Talati, Juan Manuel Zambrano Chaves, Daniel Rubin, and Imon Banerjee. “Weakly supervised language models for automated extraction of critical findings from radiology reports.” npj Digital Medicine 8, no. 1 (2025): 257.
- Das, Avisha, Amara Tariq, Felipe Batalini, Boddhisattwa Dhara, and Imon Banerjee. “Exposing Vulnerabilities in Clinical LLMs Through Data Poisoning Attacks: Case Study in Breast Cancer.” AMIA Annual Symposium; medRxiv (2024).
- Chao, Chieh-Ju, Imon Banerjee, Reza Arsanjani, Chadi Ayoub, Andrew Tseng, Jean-Benoit Delbrouck, Garvan C. Kane et al. “Evaluating large language models in echocardiography reporting: opportunities and challenges.” European Heart Journal-Digital Health 6, no. 3 (2025): 326-339.
- Tariq, Amara, Shubham Trivedi, Aisha Urooj, Gokul Ramasamy, Sam Fathizadeh, Matthew Stib, Nelly Tan, Bhavik Patel, and Imon Banerjee. “Patient-centric Summarization of Radiology Findings using Two-step Training of Large Language Models.” ACM Transactions on Computing for Healthcare 6, no. 2 (2025): 1-15.
Graph-based fusion of EHR and Radiology image
To successfully perform any clinical prediction task, it is essential to learn effective representations of various data captured during patient encounters and model their interdependencies, including patient demographics, diagnostic codes, and radiologic imaging. Graph convolutional neural networks (GCN) present an intuitive and elegant way of processing multi-modal data presented as a graph structure. We proposed novel GCN model design to understand the relationship between imaging and non-imaging data and by incorporating holistic weighted edge formation based on patient clinical history and demographic information.
AI models are currently in limited clinical rollout.
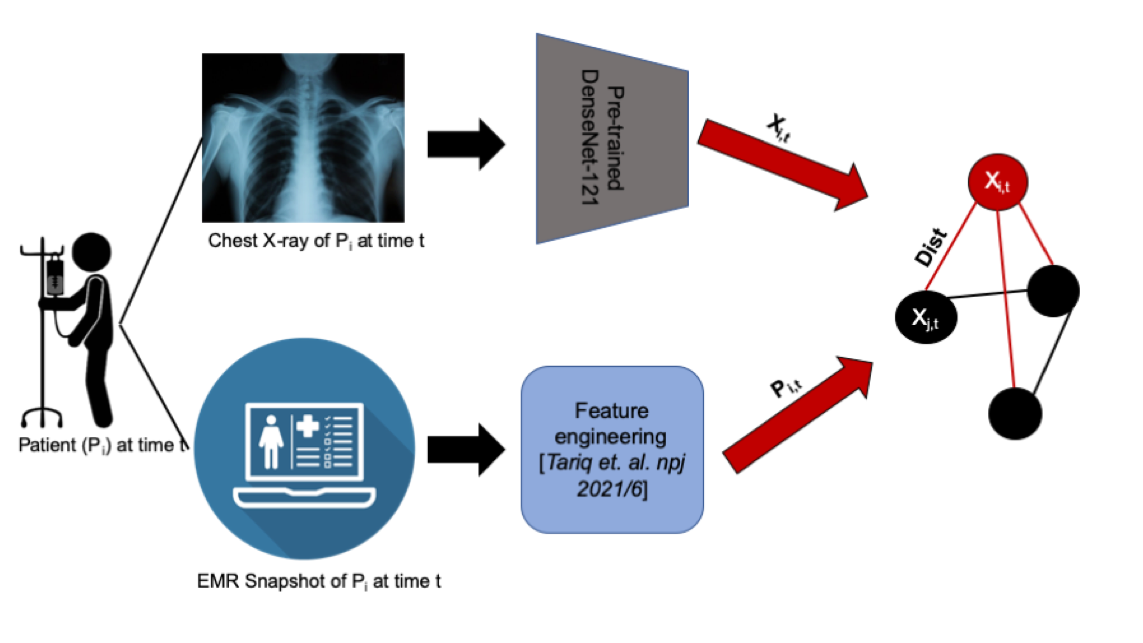
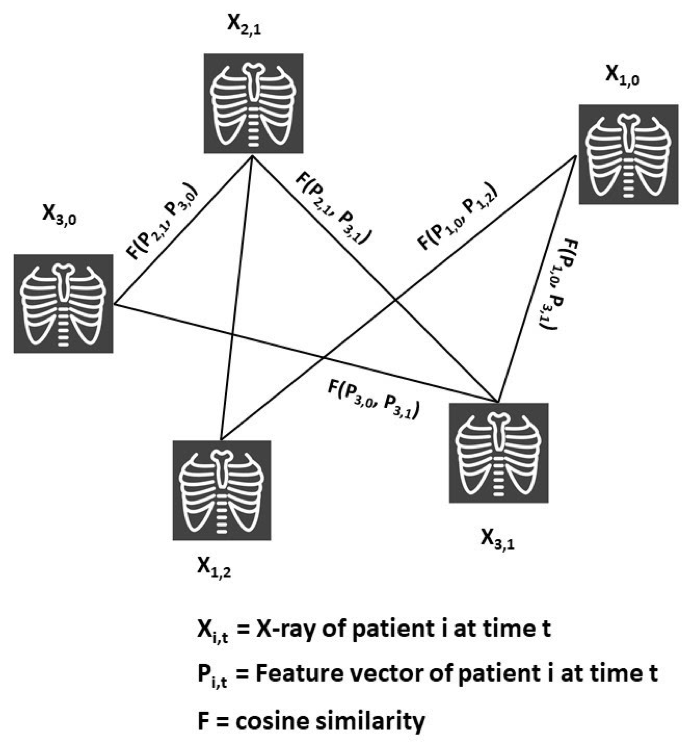

- Tariq, Amara, Lin Lancaster, Praneetha Elugunti, Eric Siebeneck, Katherine Noe, Bijan Borah, James Moriarty, Imon Banerjee, and Bhavik N. Patel. “Graph convolutional network-based fusion model to predict risk of hospital acquired infections.” Journal of the American Medical Informatics Association 30, no. 6 (2023): 1056-1067.
- Tariq, Amara, Siyi Tang, Hifza Sakhi, Leo Anthony Celi, Janice M. Newsome, Daniel L. Rubin, Hari Trivedi, Judy Wawira Gichoya, and Imon Banerjee. “Fusion of imaging and non-imaging data for disease trajectory prediction for coronavirus disease 2019 patients.” Journal of Medical Imaging 10, no. 3 (2023): 034004-034004.
- Tariq, Amara, Gurkiran Kaur, Leon Su, Judy Gichoya, Bhavik Patel, and Imon Banerjee. “Adaptable graph neural networks design to support generalizability for clinical event prediction.” Journal of biomedical informatics 163 (2025): 104794.
- Tariq, Amara, Lin Lancaster, Praneetha Elugunti, Eric Siebeneck, Katherine Noe, Bijan Borah, James Moriarty, Imon Banerjee, and Bhavik N. Patel. “Graph convolutional network-based fusion model to predict risk of hospital acquired infections.” Journal of the American Medical Informatics Association 30, no. 6 (2023): 1056-1067.
- Tang, Siyi, Amara Tariq, Jared A. Dunnmon, Umesh Sharma, Praneetha Elugunti, Daniel L. Rubin, Bhavik N. Patel, and Imon Banerjee. “Predicting 30-day all-cause hospital readmission using multimodal spatiotemporal graph neural networks.” IEEE Journal of Biomedical and Health Informatics27, no. 4 (2023): 2071-2082.
MUSE: Artificial Intelligence enabled multi-modal sensor platform for at-home health monitoring of patients with acute kidney injury (AKI)
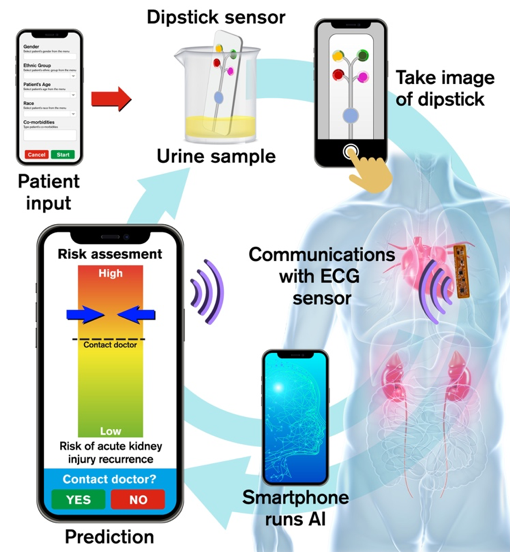
By allowing at-home AI driven self-monitoring with easy-to-use technology, MUSE can potentially identify at-risk patients without multiple clinical visits, reduce healthcare burden and improve patient outcomes through timely assessment of risks in post-AKI patients.
Aim 1. Develop a Wearable ECG patch with analog reservoir computing
Design a battery-less patch fabricated on a soft, skin-like platform to record ECG and encode the signal using an in-sensor analog reservoir computer for ultra-low energy consumption.
Aim 2. Design a Computer-vision enabled dipstick sensor platform
Develop a single, microfluidic dipstick platform with embedded colorimetric assays for detecting four markers from urine samples through automated computer vision analysis.
Aim 3. Train a Deep learning based fusion AI model
Design a deep-learning based representation learning fusion AI model that combines multimodal sensor data (urine samples and ECG) and patient medical record (past co-morbidities and demographics) for predicting risk of adverse AKI events.
Aim 4. Smart phone application development
Develop a smartphone application that will: 1) allow user to input their medical record; 2) use built-in camera to capture images of dipstick platform that detects urinary bio-markers; 3) record ECG signal using near-field communication (NFC) protocol in the smartphone; 4) run computer vision and AI algorithms on-board to process sensor data, fuse with EMR for monitoring user health, and send alerts to user.

- Bhattacharya, Amartya, Sudarsan Sadasivuni, Chieh-Ju Chao, Pradyumna Agasthi, Chadi Ayoub, David R. Holmes, Reza Arsanjani, Arindam Sanyal, and Imon Banerjee. “Multi-modal fusion model for predicting adverse cardiovascular outcome post percutaneous coronary intervention.” Physiological Measurement 43, no. 12 (2022): 124004.
- Sadasivuni, Sudarsan, Sumukh Prashant Bhanushali, Imon Banerjee, and Arindam Sanyal. “In-sensor neural network for high energy efficiency analog-to-information conversion.” Scientific reports 12, no. 1 (2022): 18253.
- Sadasivuni, Sudarsan, Monjoy Saha, Neal Bhatia, Imon Banerjee, and Arindam Sanyal. “Fusion of fully integrated analog machine learning classifier with electronic medical records for real-time prediction of sepsis onset.” Scientific reports 12, no. 1 (2022): 5711.
- Sikha, Madhu Babu, Lalith Appari, Gurudatt Nanjanagudu Ganesh, Amay Bandodkar, and Imon Banerjee. “Multi-Analyte, Swab-based Automated Wound Monitor with AI.” arXiv preprint arXiv:2506.03188 (2025).
Adversarial and Causal debiasing and adaptation — medical image case-study
Despite the expert-level performance of artificial intelligence (AI) models for various medical imaging tasks, real-world performance failures with disparate outputs for various minority subgroups limit the usefulness of AI in improving patients’ lives. AI has been shown to have a remarkable ability to detect protected attributes of age, sex, and race, while the same models demonstrate bias against historically underserved subgroups of age, sex, and race in disease diagnosis. Therefore, an AI model may take shortcut predictions from these correlations and subsequently generate an outcome that is biased toward certain subgroups even when protected attributes are not explicitly used as inputs into the model. We explore various types of bias from shortcut learning that may occur at different phases of AI model development and develop mitigation technique from preprocessing (data-centric solutions) and during model development (computational solutions) and postprocessing (recalibration of learning).
Domain adaptation – While the kidney segmentation is a required step for various clinical analyses, starting from disease detection, surgical planning, radiation dose optimization, to longitudinal characterization, the model development has largely focused on using a single-phase CT, resulting in significant performance degradation caused by drift in contrast phase appearance. We target three domain shifts to propose a robust kidney segmentation model: (i) contrast to non-contrast, (ii) arterial to venous phase, and (iii) normal functioning kidney to abnormal. Leveraging a traditional segmentation residual network (SegResNet) backbone, we design the domain adaptation for phase resiliency using an auxiliary contrast-phase classifier with confusion loss.
Causal debiasing – Our project introduces MOSCARD, a novel predictive modeling framework that integrates chest X-rays (CXR) and 12-lead electrocardiograms (ECG) to enhance cardiovascular risk assessment. While CXR provides insights into chronic conditions contributing to MACE, ECG captures real-time cardiac electrical activity and structural abnormalities. By combining these two complementary modalities, MOSCARD delivers a more holistic and accurate risk profile than traditional models based on clinical scores, CT scans, or biomarkers. Unlike much of the existing literature, which often overlooks the role of confounders and underlying causal structures in multimodal data, our approach explicitly integrates causal reasoning by incorporating comorbidities and mitigates bias.

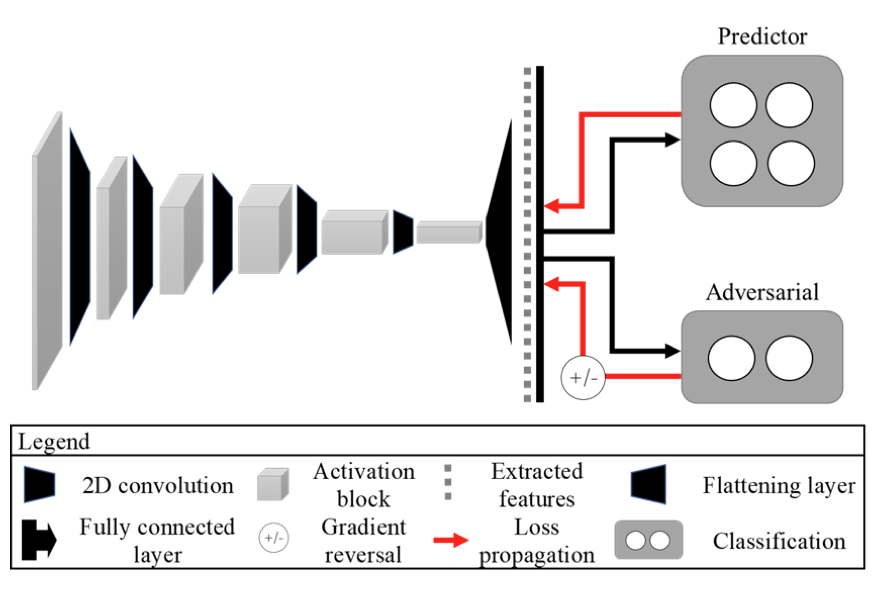
Adversarial debiasing – In our ongoing study, we develop a two-step adversarial debiasing approach with partial learning that can reduce the racial disparity while preserving the performance of the targeted task. The methodology has been evaluated onindependent medical image case-studies – skin cancer, chest X-ray, mammograms, and showed promises in bias reduction while preserving the targeted performance.
- Pi, Jialu, Juan Maria Farina, Rimita Lahiri, Jiwoong Jeong, Archana Gurudu, Hyung-Bok Park, Chieh-Ju Chao, Chadi Ayoub, Reza Arsanjani, and Imon Banerjee. “MOSCARD–Causal Reasoning and De-confounding for Multimodal Opportunistic Screening of Cardiovascular Adverse Events.” arXiv preprint arXiv:2506.19174 (2025). MICCAI 2025.
- Pi, Jialu, Juan Maria Farina, Chieh-Ju Chao, Chadi Ayoub, Reza Arsanjani, and Imon Banerjee. “Mitigating Bias in Opportunistic Screening for MACE with Causal Reasoning.” IEEE Transactions on Artificial Intelligence (2025).
- Correa-Medero, Ramon, Umar Ghaffar, Sam Fathizadeh, Bhavik Patel, Haidar Abdul-Muhsin, and Imon Banerjee. “Open-source domain adaptation to handle data shift for volumetric segmentation—use case kidney segmentation.” European Radiology (2025): 1-11.
- Banerjee, Imon, Kamanasish Bhattacharjee, John L. Burns, Hari Trivedi, Saptarshi Purkayastha, Laleh Seyyed-Kalantari, Bhavik N. Patel, Rakesh Shiradkar, and Judy Gichoya. “Shortcuts” causing bias in radiology artificial intelligence: causes, evaluation and mitigation.” Journal of the American College of Radiology (2023).
- Ramasamy, Gokul, Bhavik N. Patel, and Imon Banerjee. “Anomaly Detection using Cascade Variational Autoencoder Coupled with Zero Shot Learning.” In Medical Imaging with Deep Learning, short paper track. 2023.
- Correa-Medero, Ramon L., Bhavik Patel, and Imon Banerjee. “Adversarial Debiasing techniques towards ‘fair’skin lesion classification.” In 2023 11th International IEEE/EMBS Conference on Neural Engineering (NER), pp. 1-4. IEEE, 2023.
- Correra, Ramon, Jiwoong Jason Jeong, Bhavik Patel, Hari Trivedi, Judy W. Gichoya, and Imon Banerjee. “A robust two-step adversarial debiasing with partial learning: medical image case-studies.” In Medical Imaging 2023: Imaging Informatics for Healthcare, Research, and Applications, vol. 12469, pp. 31-38. SPIE, 2023.
- Correa-Medero, Ramón L., Rish Pai, Kingsley Ebare, Daniel D. Buchanan, Mark A. Jenkins, Amanda I. Phipps, Polly A. Newcomb et al. “Causal debiasing for unknown bias in histopathology—A colon cancer use case.” PloS one 19, no. 11 (2024): e0303415.

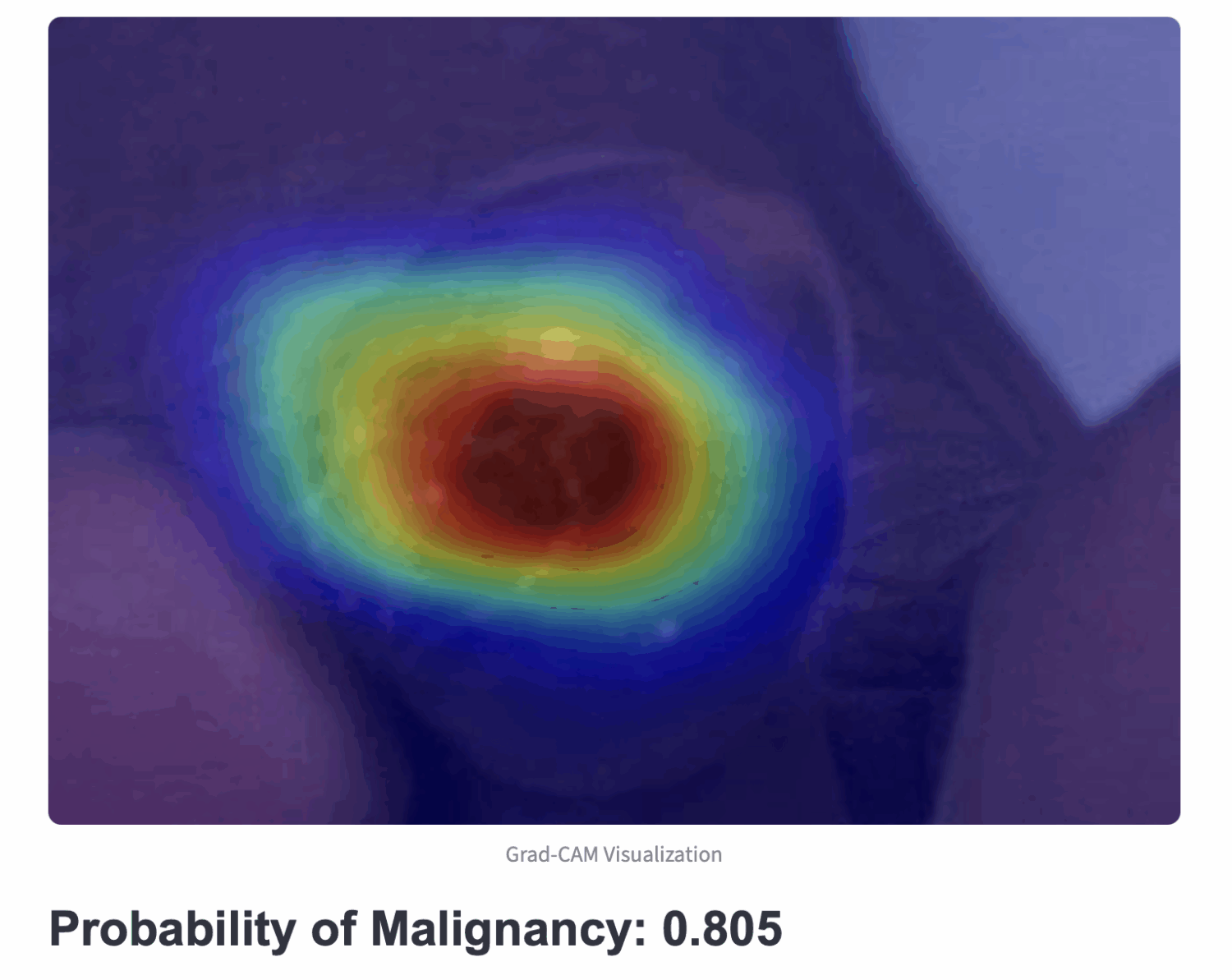
AI Technique for curating Large Cancer Databases (AI-LAD) – Breast, prostate and pancreas

Despite advances in breast cancer treatment, disparities in outcomes based on age, race/ethnicity, and socioeconomic status persist. Most existing studies are limited by small, localized datasets and focus mainly on insurance and tumor stage, lacking comprehensive long-term data. A key factor in survival—adherence to long-term treatments like endocrine therapy—is often underrepresented due to data limitations.
To address this, a large, diverse, and longitudinal breast cancer database is needed. However, manual curation is impractical due to the complexity and scale of the task. National registries like SEER only track initial treatments and lack ongoing clinical follow-up.
A multidisciplinary team is developing AI-driven natural language processing (NLP) tools to automate the extraction of long-term clinical and patient-centered outcomes from electronic medical records. Their toolset, AI-LAD, is designed to be deployed locally at institutions to parse clinical notes, pathology, and radiology reports, enabling scalable and accurate data curation for breast cancer research.
- Gupta, Anupama, Imon Banerjee, and Daniel L. Rubin. “Automatic information extraction from unstructured mammography reports using distributed semantics.” Journal of biomedical informatics 78 (2018): 78-86.
- Banerjee, Imon, Selen Bozkurt, Jennifer Lee Caswell-Jin, Allison W. Kurian, and Daniel L. Rubin. “Natural language processing approaches to detect the timeline of metastatic recurrence of breast cancer.” JCO clinical cancer informatics 3 (2019): 1-12.
- Banerjee, Imon, Kevin Li, Martin Seneviratne, Michelle Ferrari, Tina Seto, James D. Brooks, Daniel L. Rubin, and Tina Hernandez-Boussard. “Weakly supervised natural language processing for assessing patient-centered outcome following prostate cancer treatment.” JAMIA open 2, no. 1 (2019): 150-159.
- Luo, Man, Shubham Trivedi, Allison W. Kurian, Kevin Ward, Theresa HM Keegan, Daniel Rubin, and Imon Banerjee. “Automated Extraction of Patient-Centered Outcomes After Breast Cancer Treatment: An Open-Source Large Language Model–Based Toolkit.” JCO Clinical Cancer Informatics 8 (2024): e2300258.
- Chiang, Chia‐Chun, Man Luo, Gina Dumkrieger, Shubham Trivedi, Yi‐Chieh Chen, Chieh‐Ju Chao, Todd J. Schwedt, Abeed Sarker, and Imon Banerjee. “A large language model–based generative natural language processing framework fine‐tuned on clinical notes accurately extracts headache frequency from electronic health records.” Headache: The Journal of Head and Face Pain 64, no. 4 (2024): 400-409.
- Tariq, Amara, Madhu Sikha, Allison W. Kurian, Kevin Ward, Theresa HM Keegan, Daniel L. Rubin, and Imon Banerjee. “Open-Source Hybrid Large Language Model Integrated System for Extraction of Breast Cancer Treatment Pathway From Free-Text Clinical Notes.” JCO Clinical Cancer Informatics 9 (2025): e2500002.
Opportunistic screening using multi-modal data
Unstructured medical data analysis and integration of multimodal data (image + EHR) can unlock the large amount of electronic healthcare records (EHR) for clinical event prediction (e.g. ER visits, hospitalization, short-term mortality). Opportunistic screening occurs when healthcare providers use a patient’s existing clinical interactions (e.g., routine visits, imaging for unrelated issues) to screen for conditions like cancer, rather than relying solely on scheduled screening programs. Our research interest is multimodal clinical data integration and predictive modeling to perform opportunistic screening for ASCVD and pancreatic cancer. For example, A 65-year-old patient undergoes a non-gated chest CT for evaluation of chronic cough. The scan is not intended for cardiac assessment, but it incidentally a deep learning model captures the calcium in coronary arteries —a marker of subclinical atherosclerosis, and generates a patient-specific report with CAC images and guideline-based recommendations using prior clinical history.

- Design a temporal deep learning model for estimating short-term life expectancy of the patients by analyzing free-text clinical notes.
- Developed a computerized technique for assessing treatment response by analyzing noninvasive 2D/3D scans.
- Proposed a framework of data analysis tools for the automatic computation of qualitative and quantitative parameters to support effective annotation of patient-specific follow-up data.
- Developing a longitudinal machine learning approach to predict weight gain/loss in the context of insulin sensitivity and resistance by combining multiple omics data.
Papers/products
1. “Patient-specific COVID-19 resource utilization prediction using fusion AI model.” NPJ digital medicine 4, no. 1 (2021): 1-9. [link]
2. “Probabilistic Prognostic Estimates of Survival in Metastatic Cancer Patients (PPES-Met) Utilizing Free-Text Clinical Narratives.” [link]
3. “Assessing treatment response in triple-negative breast cancer from quantitative image analysis in perfusion magnetic resonance imaging.” [link]
4. “Integrative Personal Omics Profiles during Periods of Weight Gain and Loss.”, [link]
5. “Semantic annotation of 3D anatomical models to support diagnosis and follow-up analysis of musculoskeletal pathologies.” [link]
Anomaly detection for medical images
Traditional anomaly detection methods focus on detecting inter-class variations while medical image novelty identification is inherently an intra-class detection problem. For example, a machine learning model trained with normal chest X-ray and common lung abnormalities, is expected to discover and flag idiopathic pulmonary fibrosis which a rare lung disease and unseen by the model during training. The nuances from intra-class variations and lack of relevant training data in medical image analysis pose great challenges for existing anomaly detection methods. To tackle the challenges, we propose a hybrid model – nonlinear Transformation-based Embedding learn- ing for Novelty Detection (TEND). Without any out-of-distribution training data, TEND performs novelty identification by unsupervised learning of in-distribution embeddings with a vanilla AutoEncoder in the first stage and dis- criminative learning of in-distribution data and the non-linearly transformed counterparts with a binary classifier and a margin-aware objective metric in the second stage. The binary discriminator learns to distinguish the in-distribution data from the generated counterparts and outputs a class probability. The margin-aware objective is optimized jointly to include the in-distribution data in a hypersphere with a pre-defined margin and exclude the unexpected data. Even- tually, the weighted sum of class probability and the distance to margin constitutes the anomaly score.
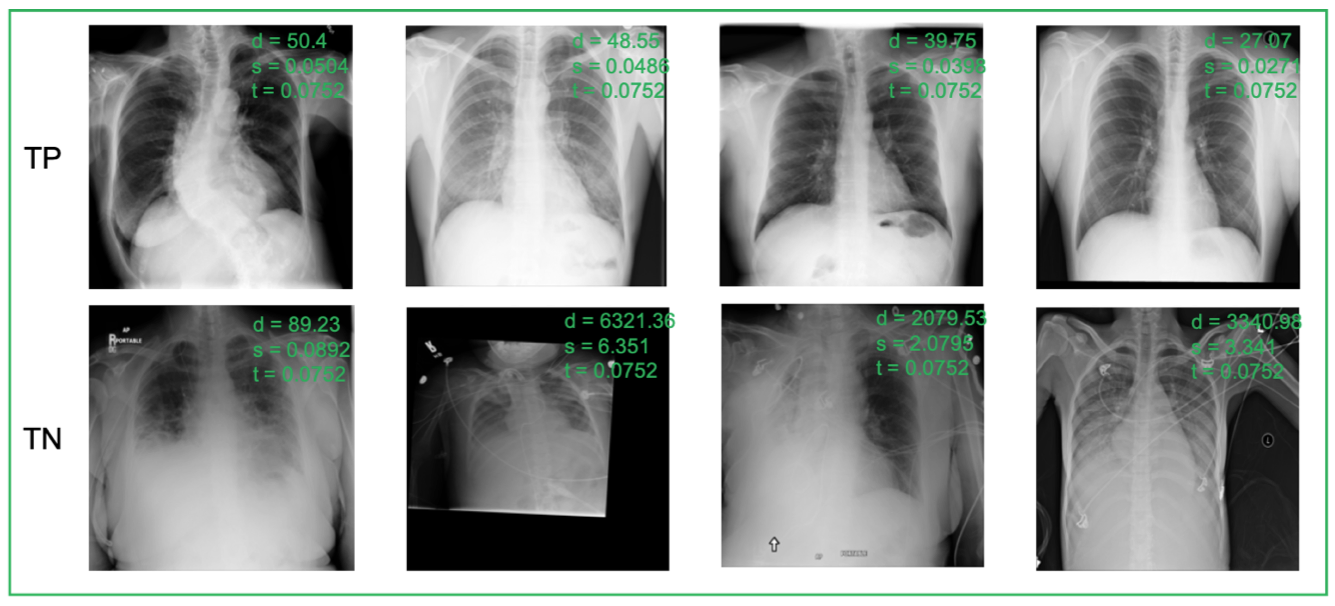
True Positive (TP, 1st row), True Negative (TN, 2nd row) predictions of TEND 500 on RSNA datasets. d: distance value from the margin learner module, p: probability outputted by the binary discriminator module, s: final score, t: optimal threshold
- Guo, Xiaoyuan, Judy Wawira Gichoya, Hari Trivedi, Saptarshi Purkayastha, and Imon Banerjee. “MedShift: Automated Identification of Shift Data for Medical Image Dataset Curation.” IEEE Journal of Biomedical and Health Informatics(2023).
- Ramasamy, Gokul, Bhavik N. Patel, and Imon Banerjee. “Anomaly Detection using Cascade Variational Autoencoder Coupled with Zero Shot Learning.” In Medical Imaging with Deep Learning, short paper track. 2023.
- Guo, Xiaoyuan, Judy Wawira Gichoya, Saptarshi Purkayastha, and Imon Banerjee. “CVAD: An Anomaly Detector for Medical Images Based on Cascade VAE.” In Workshop on Medical Image Learning with Limited and Noisy Data, pp. 187-196. Cham: Springer Nature Switzerland, 2022.
Quantitative analysis of medical images to support diagnosis.
Interested in developing computational methods that can extract quantitative information from images, integrate diverse clinical and imaging data, enable discovery of image biomarkers, and improve clinical treatment decisions. I am leading several innovative medical image analysis research projects related to cancer diagnosis, e.g. prostate cancer aggressiveness detection, histopathologic subtype classification of brain tumor, prediction of semantic features of bone tumor. I am developing a novel computational framework that can automatically interpret implicit semantic content from multimodal and/or multiparametric radiology images to enable biomedical discovery and to guide physicians in personalized care. I am responsible for the overall design of the framework, and development, execution, verification, and validation of the systems.
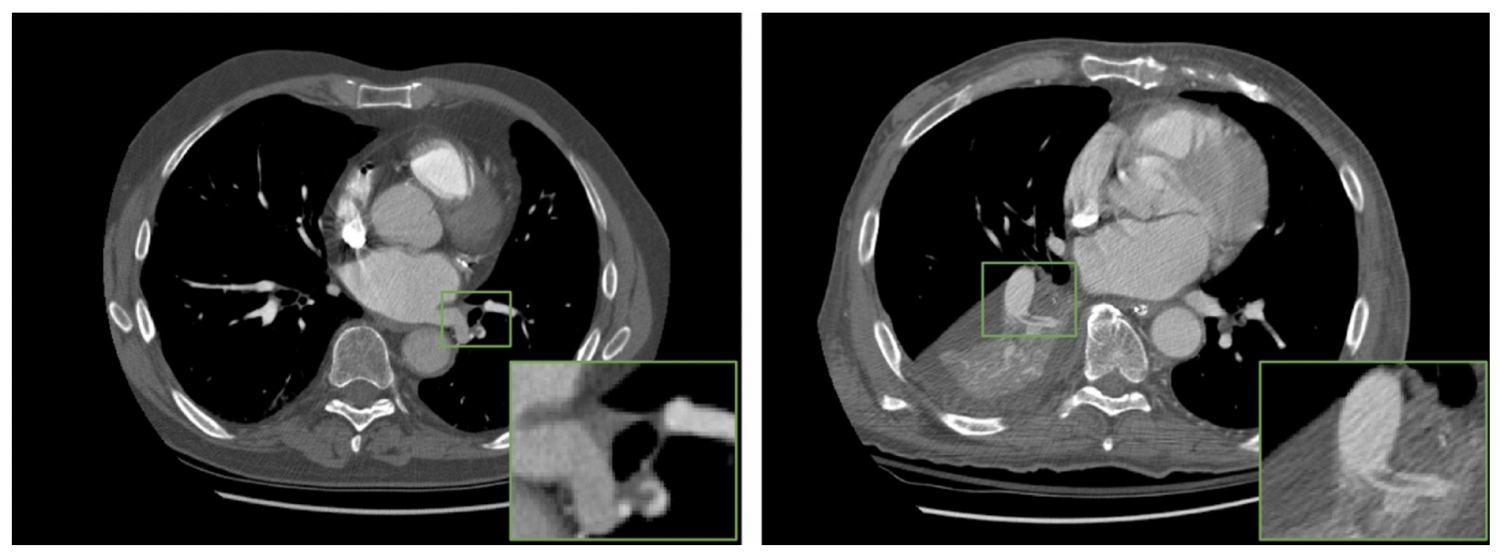
Publications and open-source code:
1. “Multimodal fusion with deep neural networks for leveraging CT imaging and electronic health record: a case-study in pulmonary embolism detection”, [link]
2. “Transfer Learning on Fused Multiparametric MR Images for Classifying Histopathological Subtypes of Rhabdomyosarcoma”, [link].
3. “Relevance feedback for enhancing content based image retrieval and automatic prediction of semantic image features: Application to bone tumor radiographs.” [link].
4. “Computerized Prediction of Radiological Observations based on Quantitative Feature Analysis: Initial Experience in Liver Lesions” [link].
5. “Computerized Multiparametric MR image Analysis for Prostate Cancer Aggressiveness-Assessment”.
Fusion of Fully Integrated Analog Machine Learning Classifier with Electronic Medical Records
The objective of this work is to develop a fusion artificial intelligence (AI) model that combines patient electronic medical record (EMR) and physiological sensor data to accurately predict early risk of sepsis and cardiovascular event. The fusion AI model has two components – an on-chip AI model that continuously analyzes patient electrocardiogram (ECG) data and a cloud AI model that combines EMR and prediction scores from on-chip AI model to predict risk score. The on-chip AI model is designed using analog circuits for sepsis prediction with high energy efficiency for integration with resource constrained wearable device. Combination of EMR and sensor physiological data improves prediction performance compared to EMR or physiological data alone, and the late fusion model has an accuracy of 93% in predicting sepsis 4 hours before onset. The key differentiation of this work over existing sepsis prediction literature is the use of single modality patient vital (ECG) and simple demographic information, instead of comprehensive laboratory test results and multiple vital signs. Such simple configuration and high accuracy makes our solution favorable for real-time, at-home use for self-monitoring.
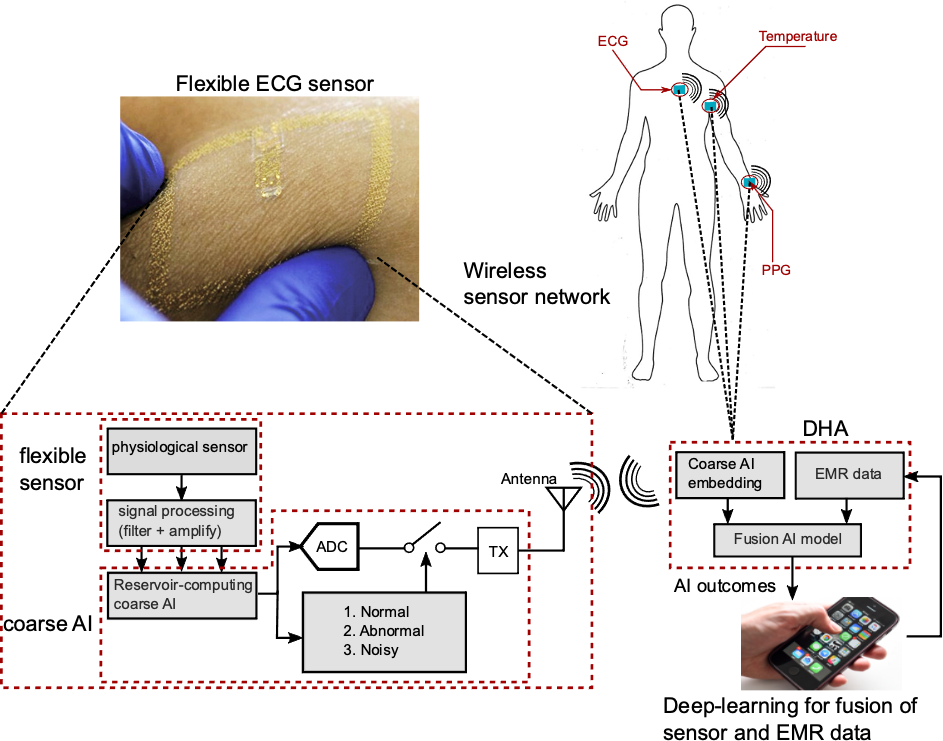
Publications and open-source code:
1. “Recurrent Neural Network Circuit for Automated Detection of Atrial Fibrillation from Raw ECG.” In 2021 IEEE International Symposium on Circuits and Systems (ISCAS), 2021. [link]
2. “Fully Integrated Analog Machine Learning Classifier Using Custom Activation Function for Low Resolution Image Classification,” [link][code]
3. “Digital Machine Learning Circuit for Real-Time Stress Detection from Wearable ECG Sensor.” [link][code]
Natural Language Processing on clinical notes
The lack of labeled data creates a data “bottleneck” for developing deep learning models for medical imaging. However, healthcare institutions have millions of imaging studies which are associated with unstructured free text radiology reports that describe imaging features and diagnoses, but there are no reliable methods for leveraging these reports to create structured labels for training deep learning models. Unstructured free text thwarts machine understanding, due to the ambiguity and variations in language among radiologists and healthcare organizations.
My research is focused in developing methods to extract structured annotations of medical images from radiology reports for training complex deep learning models.
Our method has outperformed many existing NLP algorithms on several radiology report annotation tasks (CT reports, mammography reports, US reports, and X-ray reports), as well as can infer targeted information from heterogeneous clinical notes (e.g., hospital notes, discharge summary, progress notes).
Publications and open-source code:
1. “Weakly supervised temporal model for prediction of breast cancer distant recurrence.” [link][code]
2. “Radiology Report Annotation using Intelligent Word Embeddings: Applied to Multi-institutional Chest CT Cohort,” [link][code]
3. “Weakly supervised natural language processing for assessing patient-centered outcome following prostate cancer treatment.” [link][code]
4. “Comparative effectiveness of convolutional neural network (CNN) and recurrent neural network (RNN) architectures for radiology text report classification.” [link][code available upon request]
5. A Scalable Machine Learning Approach for Inferring Probabilistic US-LI-RADS Categorization [link][code available upon request]
6. “Development and Use of Natural Language Processing for Identification of Distant Cancer Recurrence and Sites of Distant Recurrence Using Unstructured Electronic Health Record Data.” [link]